Events
- 1
- 2
Note, this map only displays events that have geolocation information in
TeSS.
For the complete list of events in TeSS, click the grid tab.
TeSS makes use of some necessary cookies to provide its core functionality. Additionally, we make use of Google Analytics to discover how people are using TeSS in order to help us improve the service. To opt out of this, choose the "Allow necessary cookies" option.
See our Privacy Policy for more information.
You can modify your cookie preferences at any time here, or from the link in the footer.
Scientific topics: Bioinformatics
In the left-hand column of the Google Calendar main view, click the arrow to the right of "Other calendars" and click "Add by URL". In the form that appears, paste in the URL from the box above, and click the button to confirm.
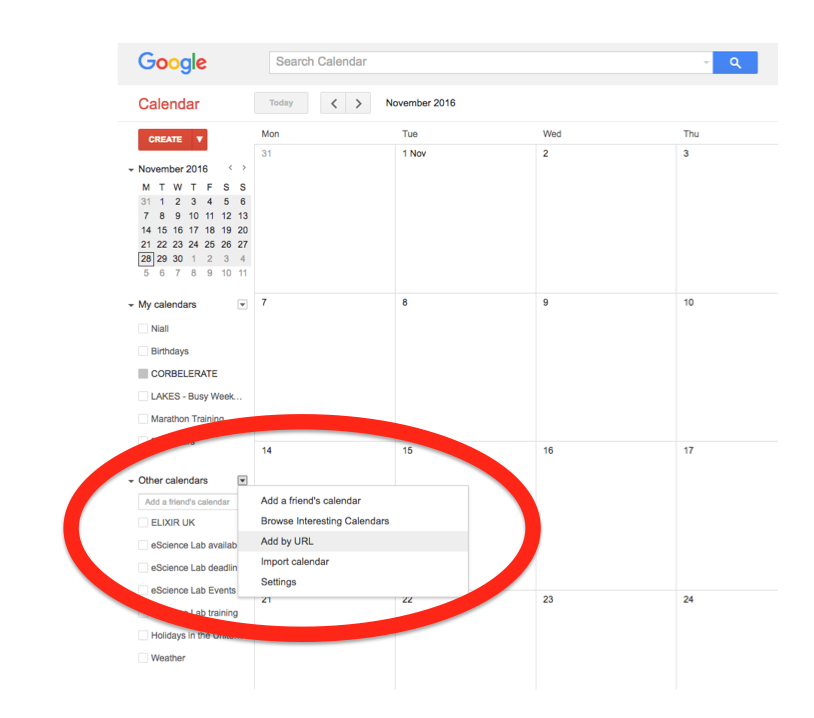
Please note, it may take a while for newly created events in TeSS to synchronise with your Google Calendar.
16 - 20 May 2024
Cambridge, United Kingdom
Face-to-face
Bioinformatics Data mining Data visualisation Functional genomics Transcriptomics HDRUK17 May 2024 @ 09:00 - 12:00
Tallinn, Estonia
Face-to-face
Bioinformatics g:Profiler24 May 2024 @ 08:30 - 16:30
Cambridge, United Kingdom
Face-to-face
Bioinformatics Data mining Data visualisation Phylogenetics HDRUK24 May 2024 @ 14:00 - 17:00
Online
Bioinformatics g:Profiler18 - 19 June 2024
Cambridge, United Kingdom
Face-to-face
Bioinformatics Data visualisation Metabolomics HDRUK20 June 2024 @ 08:30 - 12:00
Cambridge, United Kingdom
Face-to-face
Bioinformatics Data visualisation Data mining Sequencing HDRUK21 - 25 June 2024
Cambridge, United Kingdom
Face-to-face
Bioinformatics Functional genomics Data visualisation Transcriptomics Data mining HDRUK26 - 27 June 2024
Cambridge, United Kingdom
Face-to-face
Bioinformatics Biology Data visualisation Proteomics HDRUK5 July 2024 @ 08:30 - 16:00
Cambridge, United Kingdom
Face-to-face
Bioinformatics Functional genomics Data visualisation ChIP-seq Data mining HDRUK18 July 2024 @ 08:30 - 16:30
Cambridge, United Kingdom
Face-to-face
Bioinformatics Biology HDRUK
Note, this map only displays events that have geolocation information in
TeSS.
For the complete list of events in TeSS, click the grid tab.